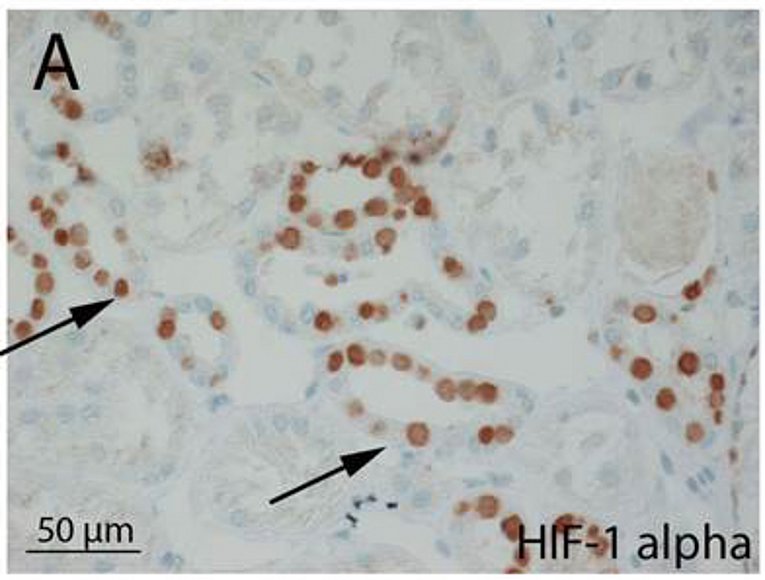
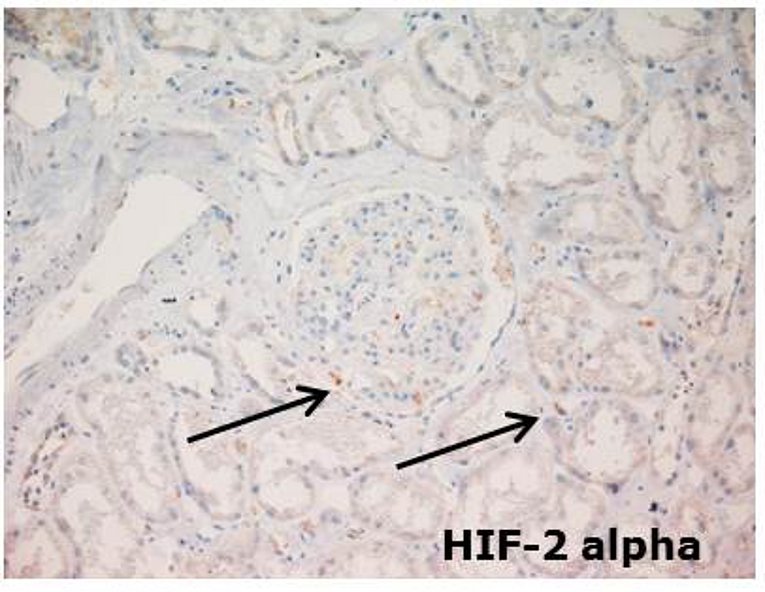
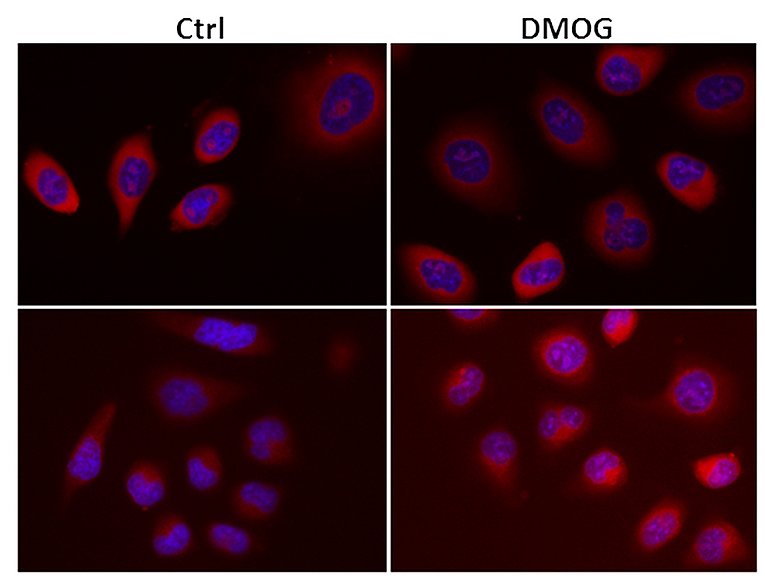
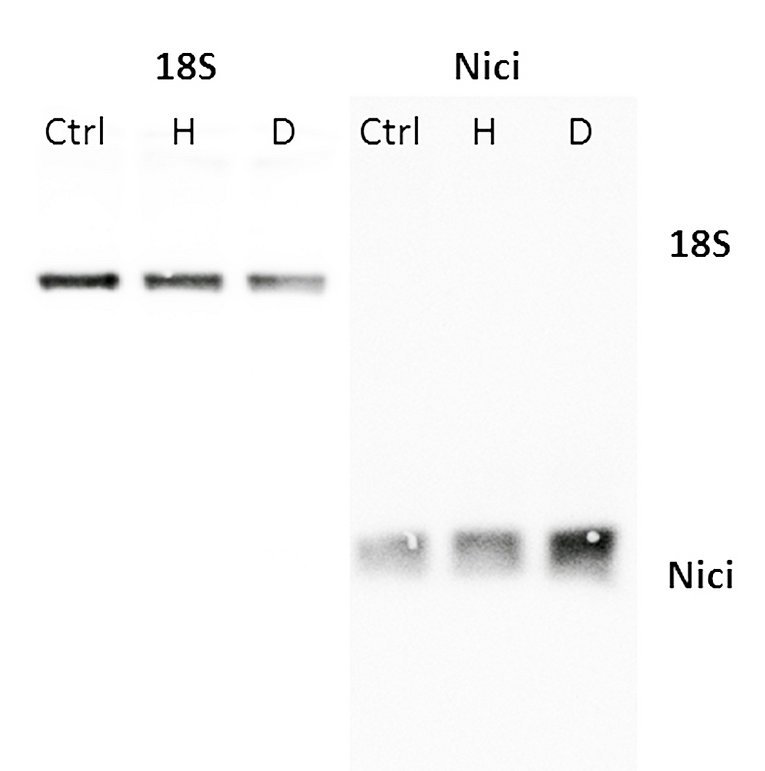
Hypoxia plays an important role in many physiologic and pathophysiologic processes. Hypoxia-inducible transcription factors (HIF) are central to hypoxic gene regulation. By binding DNA at hypoxia responsive elements HIFs activate transcription of genes such as erythropoietin, VEGFA, Cyclin D1 and glycolytic enzymes that regulate adaptive processes in the cell and the whole organism.
HIFs can activate protective mechanisms which lead to a better tubular function in acute kidney injury. In contrast, in VHL-associated renal cancer permanent activation of HIFs can contribute to disease progression.
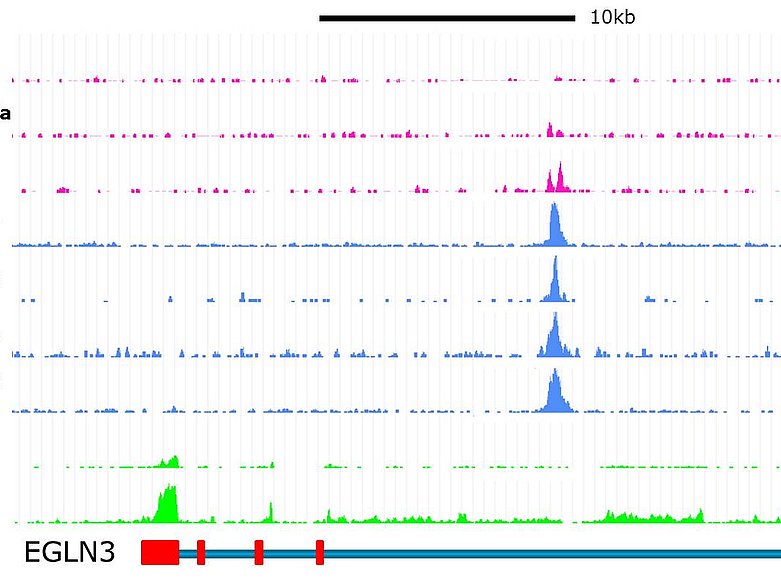
We aim to identify cell type specific functions and epigenetic characteristics of HIF-DNA binding as well as mechanisms that directly influence HIF-DNA interactions. We use modern technologies to identify HIF-binding sites and analyse protein-DNA interactions (Next generation sequencing: ChIP-seq, FAIRE-seq, RNA-seq). We evaluate the transactivation potential of these sites by genome-editing (CRISPR/Cas, TALEN) in relevant cell lines. Furthermore, we explore interactions of HIF-binding, epigenetics and single nucleotide polymorphisms in the context of renal cancer and acute and chronic kidney disease.
Collaborations
- Prof. Sir Peter Ratcliffe, Prof. David Mole, Centre for Cellular and Molecular Physiology, Oxford, UK
- Prof. Günther Greiner, Lehrstuhl für Informatik 9, Friedrich-Alexander-Universität Erlangen-Nürnberg
- Prof. Arndt Hartmann, Dr. Christine Stöhr, Institut für Pathologie, Universitätsklinikum Erlangen
- SFB 1350, Universität Regensburg